CS194-26 Project 3 Face Morphing
Weichao Chen
PART I - Manual Image Stitching
Homography Matrix Calculation
We need four pairs of corresponding points between two images to uniquely identify the projective homography matrix by solving the following system of equations. (8 unknows and equations)
Once we have the matrix , we can apply it on the project an image onto the reference surface by having . More on this on StackExchange
Image Rectifying
Below are examples that shows projective transformation in use.
Flyer

Screen

Image Stiching
By taking two pictures with different point of views but overlapped field of view, say pic1 and pic2. Then warp pic1 using the homography matrix defined between the two pictures.
Flyer

Shelf
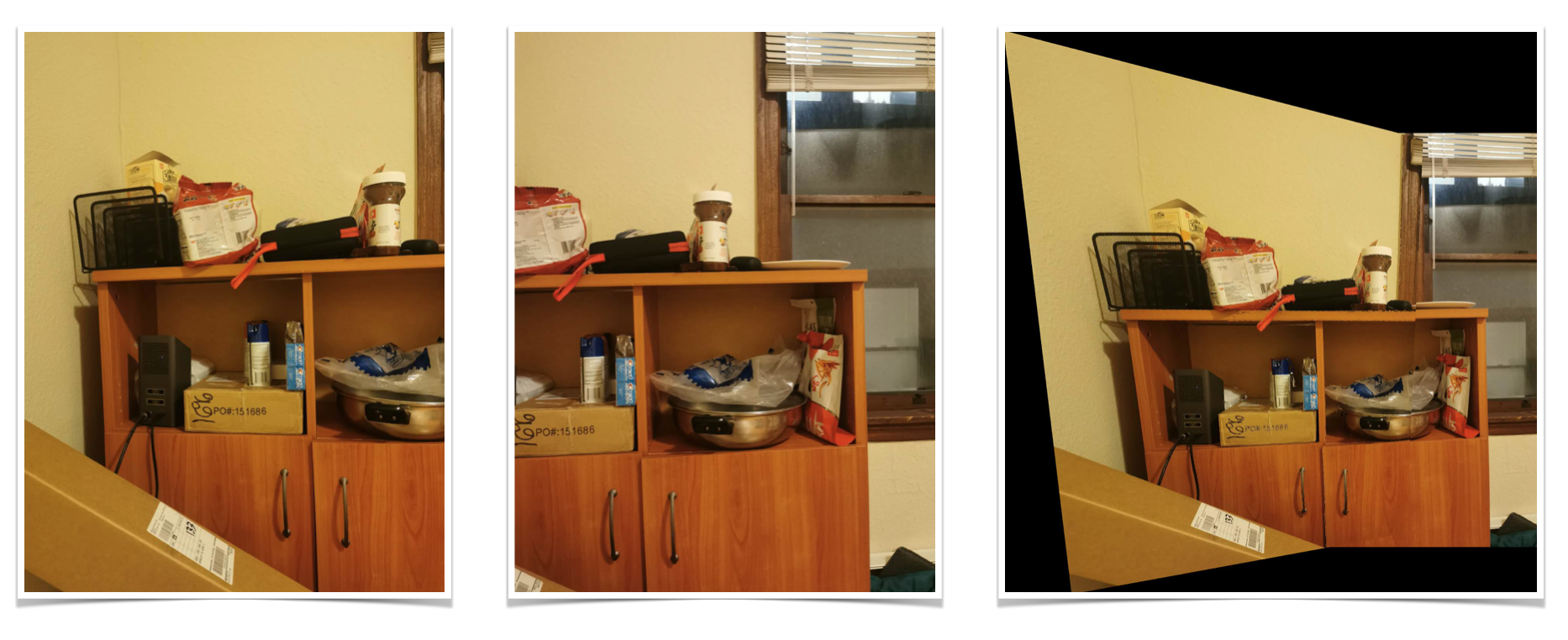

Kitchen
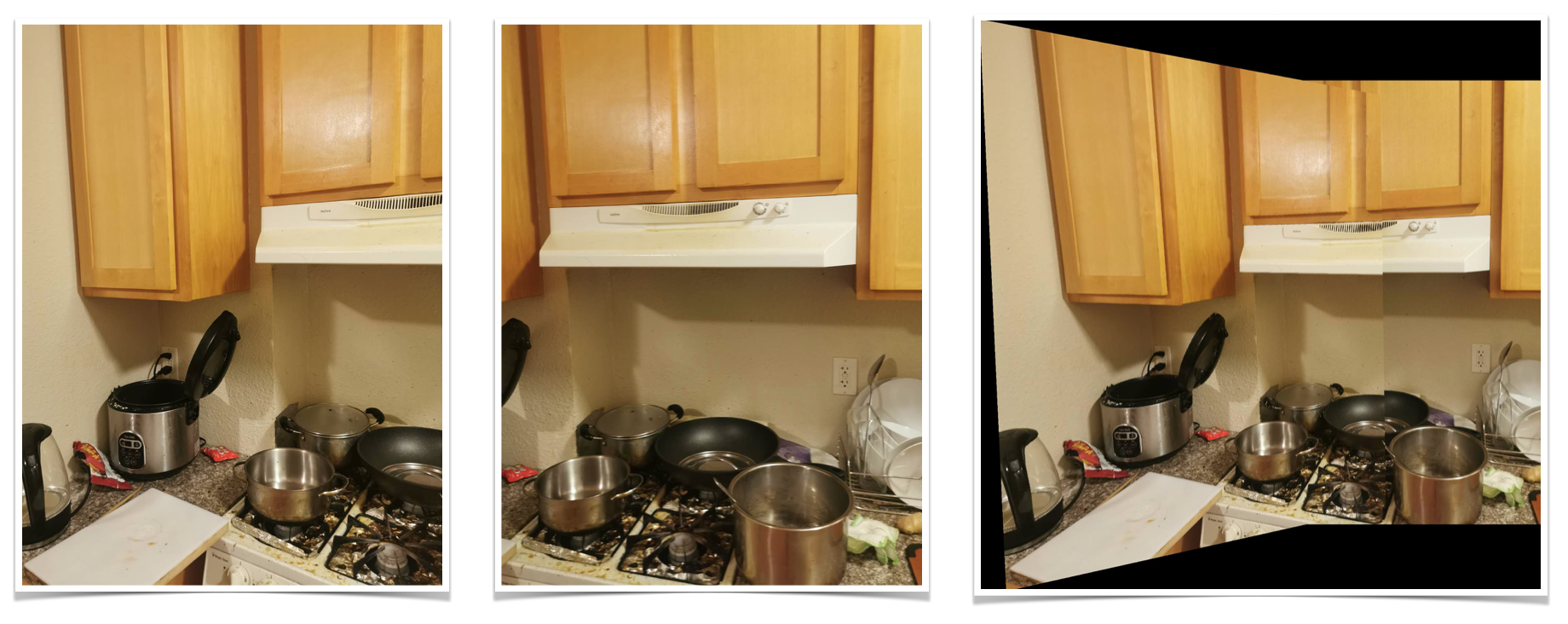

PART II - Automatic Image Stitching
In this part we attempt to automate the image stiching process by using
- HarrisCorner algorithm to compute the "cornerness" of each pixel
- Adaptive Non-Maximum Suppression to select a handful of pixels with the highest level of "corerness"
- Patch Texture Feature Extraction for each selected candidate corner coordinate
- SSD as metrics to compare and define correspondence relationship between the corner coordinates for two images
- RANSAC to select best correspondence used for computing , the homography matrix
Harris Corner and ANMS
After Harris Corner computes the cornerness score at each coordinate, we need to select a handful of pixel with most "corner-like" charateristics. If we simply use the top k coordinates with the higest scores, we might end up with points cluster together which with make the homography matrix less robust. ANMS on the other hand, by computing the suppression radius for each point i.e. the distance until the closest point has a higher "cornerness" score, ensures that the selected pixels are more evenly distributed across the image.
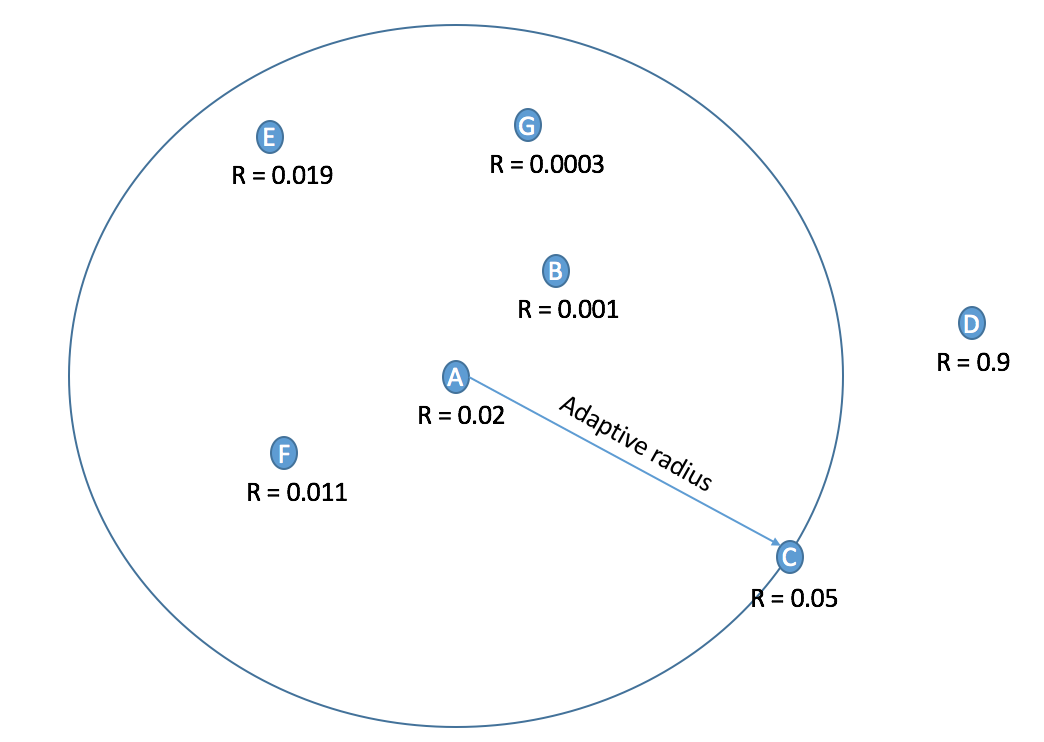
Photo Credit: Anisha Gartia, Georgia Tech
Example of points selected by ANMS at
Patch Texture Feature Extraction
Next up we'd calculate a high-dimensional feature descriptor at each point so we can defind correspondence between points across different images. Ideally, these descriptor should be invariant to rotation and scaling. For the sake of simplicity though, I use a 40 * 40 window to extract the gradient sub-region containing each point, down-sample such window to a 8 * 8 window (patch), convolve it against a Gaussian filter so that points close-up gets a higher weight, and then normalize to have a standard deviation of 1 and mean of 0 before I flatten the matrix into a 1 * 64 vector.
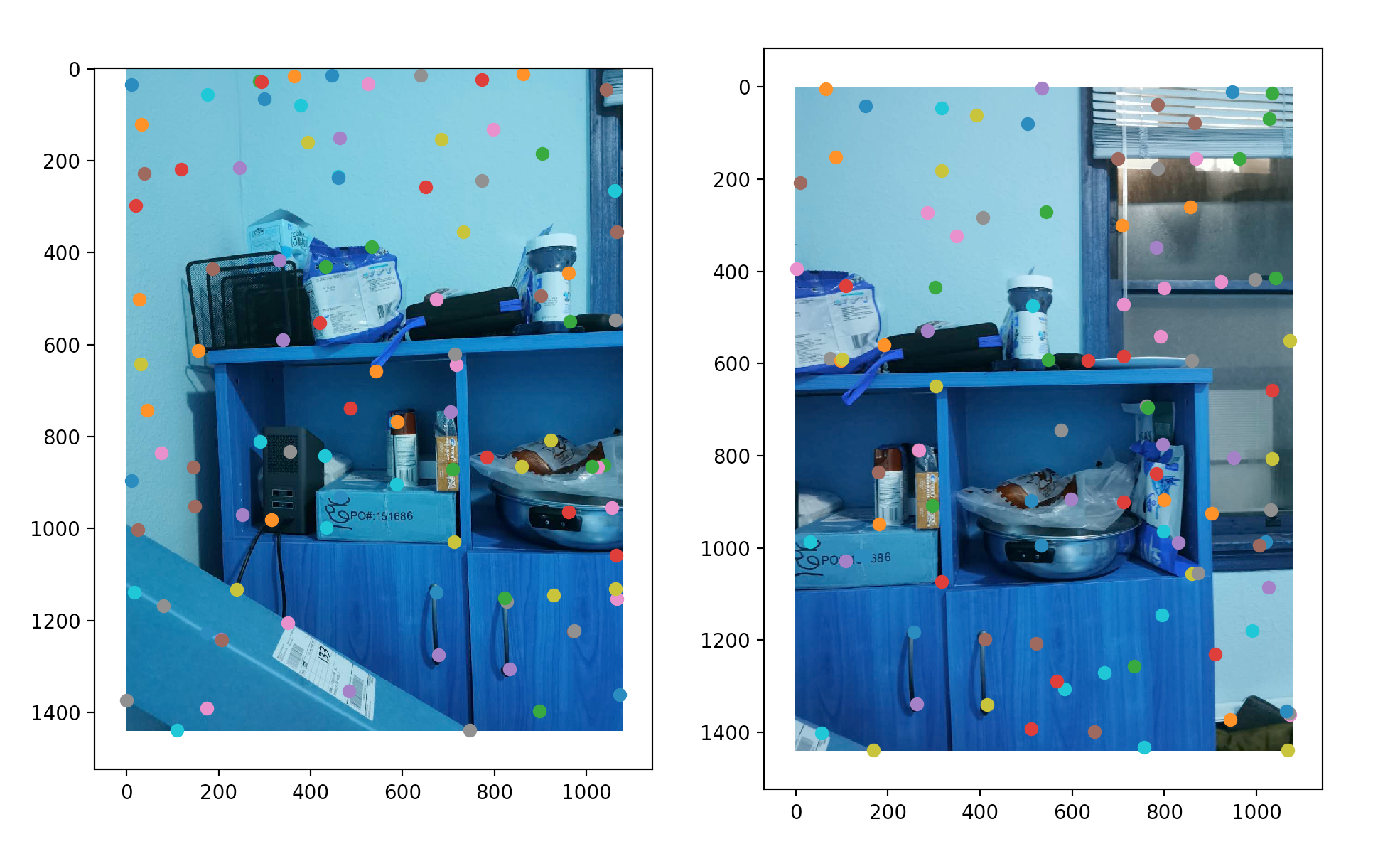
Defining Point Correspondences
Now that we have k points from two different images and the descriptor vector v associated with them, we simply calculate the pair-wise Euclidean distance using v and match the two points with the shortest distance. After that, we select the pairs where the distance is below a threshold.
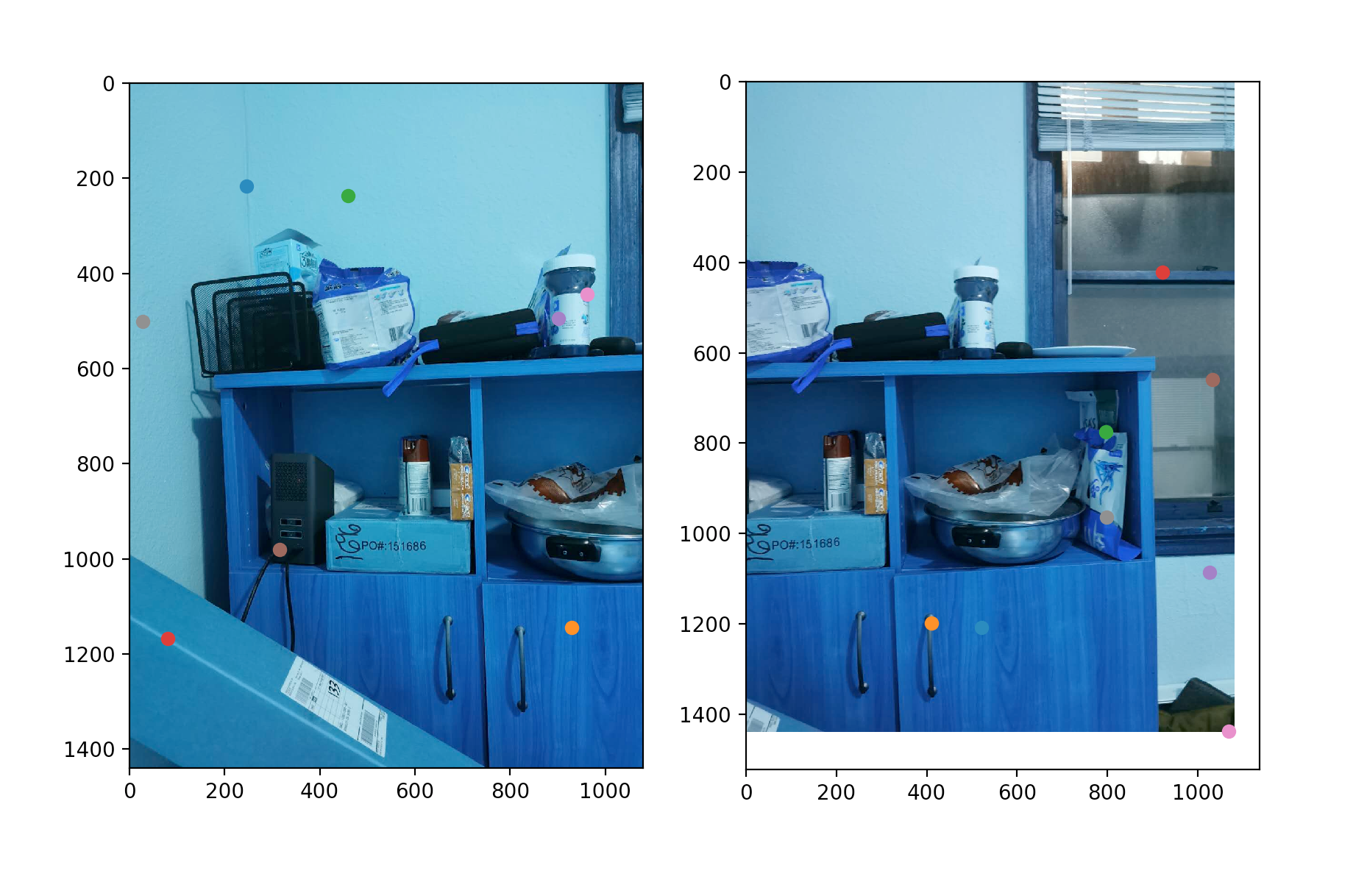
RANSAC
While we root for the matching algorithm to find correct correspondence, incorrect matches still exist. Yet, it majority of the matches are correct, we can use the RANSAC consept to find the dominant structure and ignore the outliers. Basically, we randomly select 8~12 points, calculate the homography matrix H. Using the rest of the correspondence points , we calculate the projected location using the homography matrix , and compare that with . If the distance is below a predefined threshold, we could the point as an inlier.

Photo Credit: Mehrdad Heydarzadeh
Mosaic Examples
Finally, here're some example using the stitching procedure
| shelf_left | shelf_right | shelf_merged |
|---|---|---|
 |  |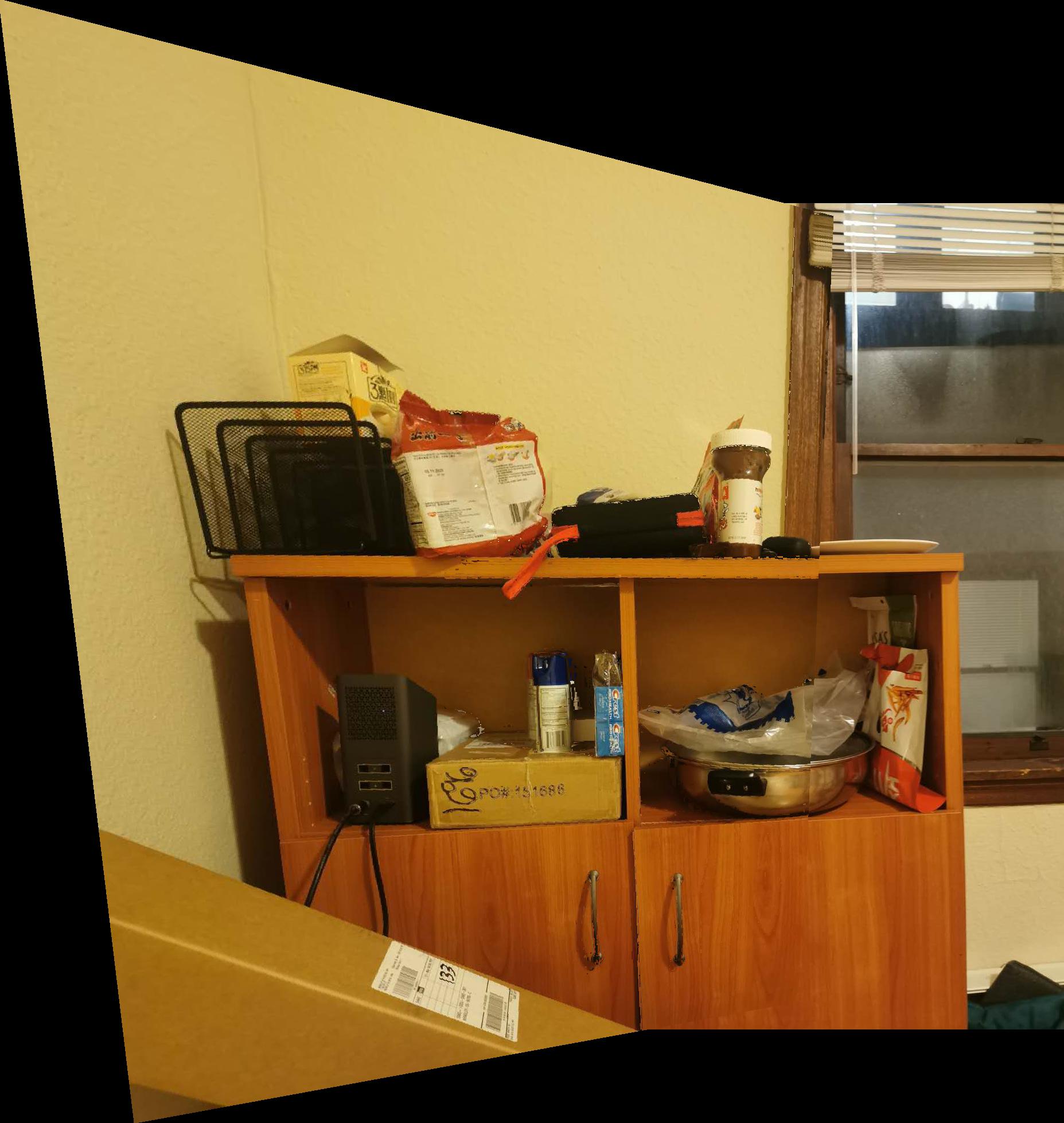  |
| kitchen_left | kitchen_right | kitchen_merged |
|---|---|---|
 |  |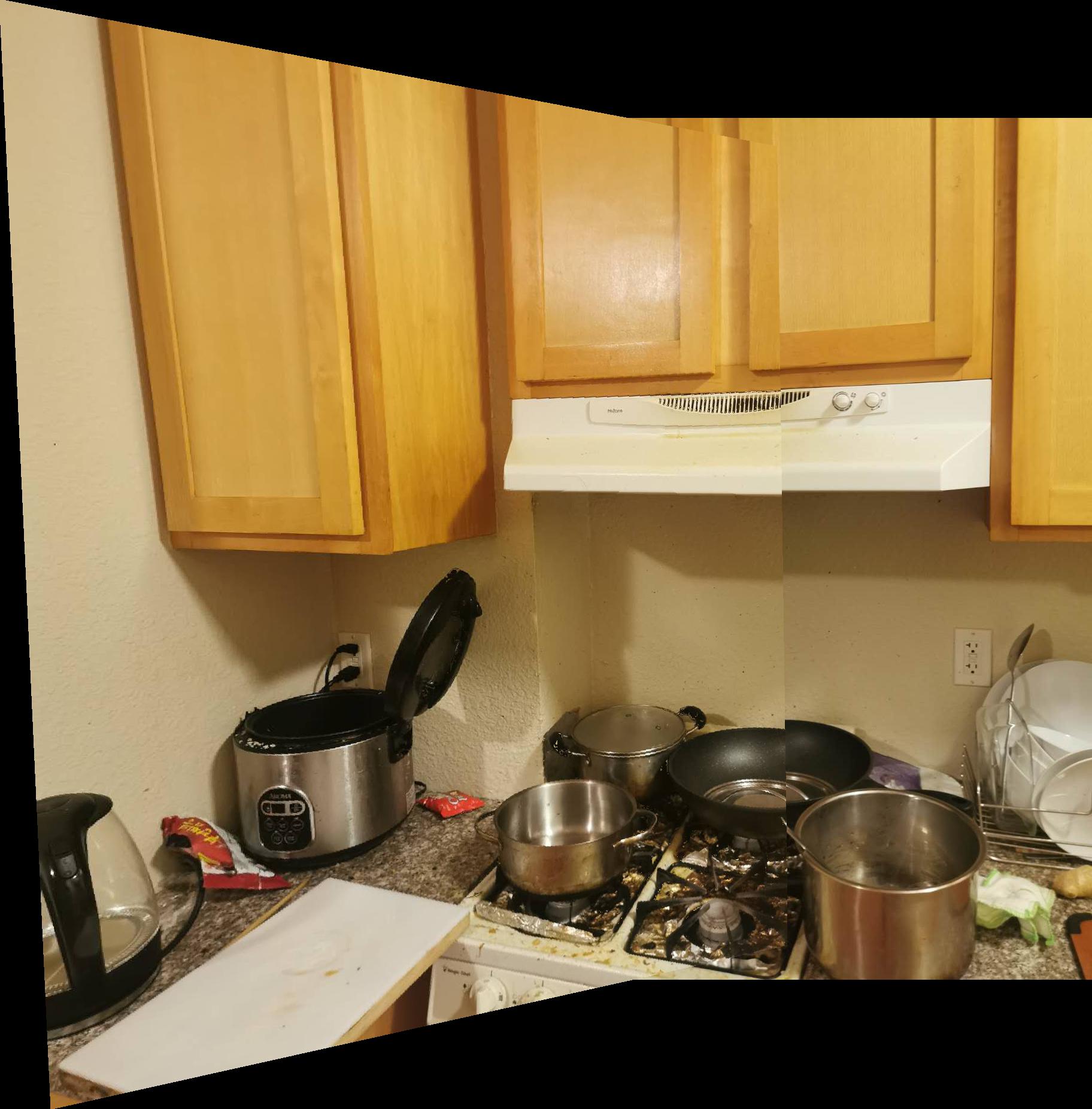  |
| trash_left | trash_right | trash_merged |
|---|---|---|
 |  |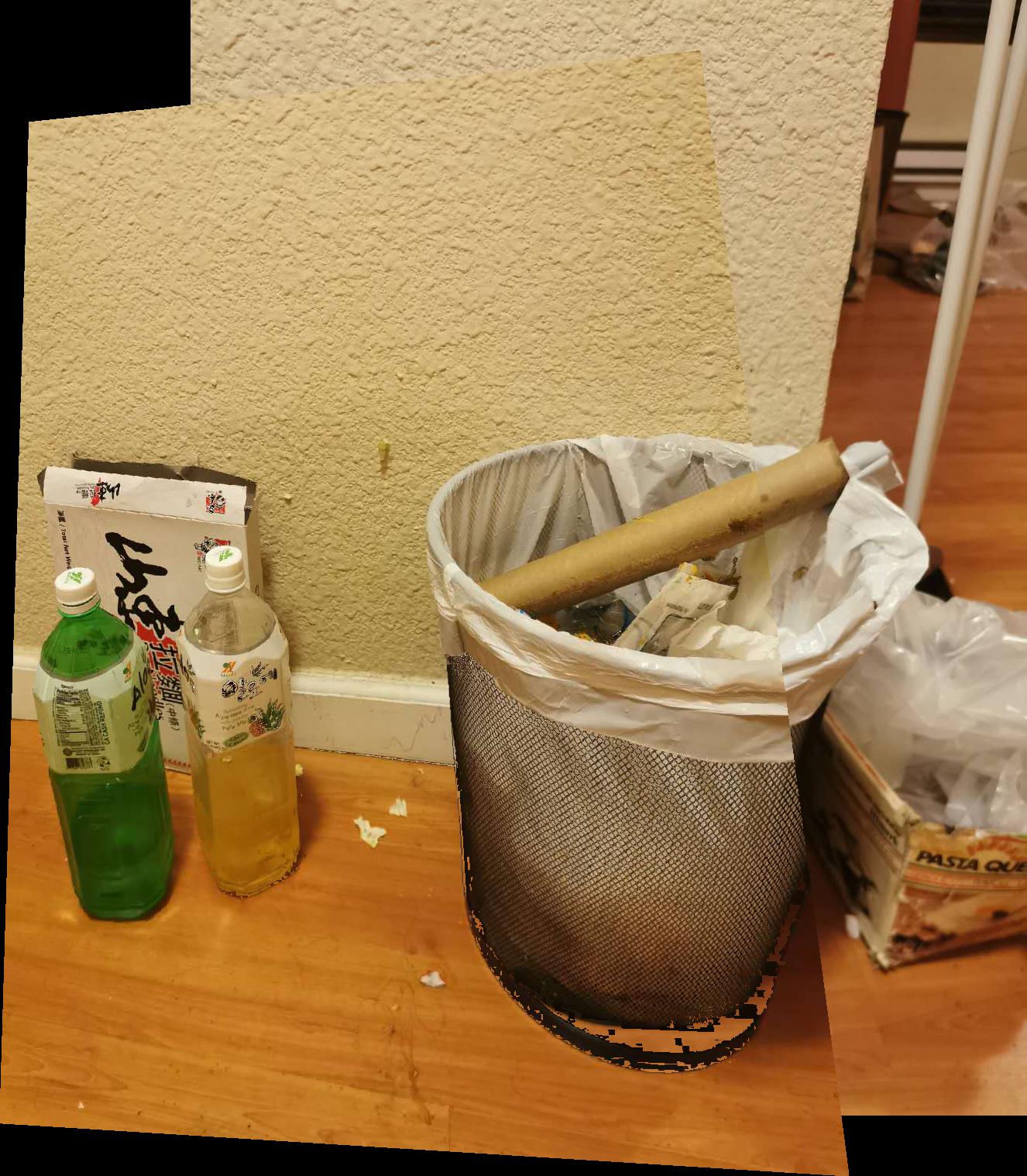  |
What I Learn
The intuition behind RANSAC is pretty simple, although the name is pretty intimidating. Moving forward, I'll be interested in understanding finding scale, rotation invariance feature descriptor